statistics-surface - Surface-based mass-univariate analysis with SurfStat¶
This command performs statistical analysis (e.g. group comparison, correlation) on surface-based features using the general linear model (GLM). To that aim, the pipeline relies on the Matlab toolbox SurfStat designed for statistical analyses of univariate and multivariate surface and volumetric data using the GLM [Worsley et al., 2009].
Surface-based measurements are analyzed on the FsAverage surface template (from FreeSurfer).
Currently, this pipeline can handle cortical thickness measurements from T1 images t1-freesurfer pipeline or map of activity from PET data using pet-surface pipeline.
Note
We are aware that the SurfStat toolbox is not maintained anymore. The reasons why we rely on it are: 1) its great flexibility; 2) our profound admiration for the late Keith Worsley.
Prerequisites¶
You need to process your data with the t1-freesurfer pipeline for measurements of cortical thickness measurements from T1 images or pet-surface pipeline for measurements of activity map from PET.
Do not hesitate to have a look at the paragraph Specifying what surface data to use if you want to use your own surface feature.
Dependencies¶
If you only installed the core of Clinica, this pipeline needs the installation of Matlab and FreeSurfer 6.0 on your computer.
You can find how to install these software packages on the third-party page.
Note that the Matlab Statistics and Machine Learning Toolbox is required.
Compatibility issue with Matlab R2019 / R2020
It has been reported that newer versions of Matlab (see details on GitHub) were not compatible with this pipeline. For the moment, we advise you to use at most the R2018b version. Matlab versions between 2015 and 2018 are known to work.
Running the pipeline¶
The pipeline can be run with the following command line:
clinica run statistics-surface [OPTIONS] CAPS_DIRECTORY GROUP_LABEL {t1-freesurfer|pet-surface|custom-pipeline}
{group_comparison|correlation} SUBJECT_VISITS_WITH_COVARIATES_TSV CONTRAST
where:
CAPS_DIRECTORYis the folder containing the results of thet1-freesurferorpet-surfacepipeline and the output of the present command, both in a CAPS hierarchy.GROUP_LABELis a string defining the group label for the current analysis, which helps you keep track of different analyses.- The third positional argument defines the type of surface-based feature: it can be
t1-freesurferfor cortical thickness,pet-surfacefor projected PET data orcustom-pipelinefor you own data in CAPS directory (see below for details). - The fourth positional argument is a string defining the type of analysis of your model, choose one between
group_comparisonandcorrelation. SUBJECT_VISITS_WITH_COVARIATES_TSVis a TSV file containing a list of subjects with their sessions and all the covariates and factors in your model (the content of the file is explained in the Example subsection).CONTRASTis a string defining the contrast matrix or the variable of interest for the GLM, e.g.grouporage.
Pipeline options:
-c, or--covariates: Covariates must be provided one at the time:-c covariate_1 -c covariate_2, or equivalently--covariates covariate_1 --covariates covariate_2. Each covariate must match the name of the TSV file. By default, no covariate is considered.--full_width_at_half_maximum: FWHM for the surface smoothing. Default value is20.
Pipeline options if you use inputs from the pet-surface pipeline:
--acq_label: Name of the label given to the PET acquisition, specifying the tracer used (trc-<acq_label>).--suvr_reference_region: Reference region used to perform intensity normalization (i.e. dividing each voxel of the image by the average uptake in this region) resulting in a standardized uptake value ratio (SUVR) map. It can becerebellumPons(used for amyloid tracers) orpons(used for FDG).
Tip
Check the Example subsection for further clarification.
Outputs¶
Group comparison analysis¶
Results are stored in the following folder of the CAPS hierarchy: groups/<group_id>/statistics/surfstat_group_comparison/.
The main outputs for the group comparison are:
<group_id>_<group_1>-lt-<group_2>_measure-<label>_fwhm-<label>_correctedPValue.jpg: contains both the cluster level and the vertex level corrected p-value maps, based on the random field theory.<group_id>_<group_1>-lt-<group_2>_measure-<label>_fwhm-<label>_FDR.jpg: contains corrected p-value maps, based on the false discovery rate (FDR).<group_id>_participants.tsvis a copy of thesubject_visits_with_covariates_tsvparameter file.<group_id>_glm.jsonis a JSON file containing all the model information of the analysis (i.e. what you wrote on the command line).
The <group_1>-lt-<group_2> means that the tested hypothesis is: "the measurement of <group_1> is lower than (lt) the measurement of <group_2>".
The pipeline includes both contrasts so *<group_2>-lt-<group_1>* files are also saved.
The FWHM value corresponds to the size in mm of the kernel used to smooth the surface and can be 5, 10, 15, 20.
Analysis with cortical thickness (respectively PET data) will be saved under the _measure-ct keyword (respectively the _measure-<acq_label> keyword).
Tip
See the Example subsection for further clarification.
Correlations analysis¶
Results are stored in the following folder of the
CAPS hierarchy:
groups/<group_id>/statistics/surfstat_correlation/.
The main outputs for the correlation are:
<group_id>_correlation-<label>_contrast-<label>_measure-<label>_fwhm-<label>_correctedPValue.jpg: contains p-value maps corrected at both the cluster and vertex levels, based on the random field theory.<group_id>_correlation-<label>_contrast-<label>_measure-<label>_fwhm-<label>_FDR.jpg: contains corrected p-value maps, based on the false discovery rate (FDR).<group_id>_correlation-<label>_contrast-<label>_measure-<label>_fwhm-<label>_T-statistics.jpg: contains the maps of T statistics.<group_id>_correlation-<label>_contrast-<label>_measure-<label>_fwhm-<label>_Uncorrected p-value.jpg: contains the maps of uncorrected p-values.<group_id>_participants.tsvis a copy of thesubject_visits_with_covariates_tsvparameter file.<group_id>_glm.jsonis a JSON file summarizing the parameters of the analysis (i.e. what you wrote on the command line).
The correlation-<label> describes the factor of the model, which can be for example age.
The contrast-<label> is the sign of your factor which can be negative or positive.
Analysis with cortical thickness (respectively PET data) will be saved under the _measure-ct keyword (respectively the _measure-<acq_label> keyword).
Note
The full list of output files can be found in the ClinicA Processed Structure (CAPS) specifications.
Example¶
Comparison analysis¶
Let's assume that you want to perform a group comparison of cortical thickness between patients with Alzheimer’s disease (group_1 will be called AD) and healthy subjects (group_2 will be called HC).
ADvsHC will define the group_label.
The TSV file containing the participants and covariates will look like this:
participant_id session_id sex group age
sub-CLNC0001 ses-M000 Female CN 71.1
sub-CLNC0002 ses-M000 Male CN 81.3
sub-CLNC0003 ses-M000 Male CN 75.4
sub-CLNC0004 ses-M000 Female CN 73.9
sub-CLNC0005 ses-M000 Female AD 64.1
sub-CLNC0006 ses-M000 Male AD 80.1
sub-CLNC0007 ses-M000 Male AD 78.3
sub-CLNC0008 ses-M000 Female AD 73.2
Note that to make the display clearer, the rows contain successive tabs, which should not happen in an actual TSV file.
We call this file ADvsHC_participants.tsv.
For this group comparison, we will use age and sex as covariates.
As a result, the command line will be:
clinica run statistics-surface caps_directory ADvsHC t1-freesurfer group_comparison ADvsHC_participants.tsv group -c age -c sex
The parameters of the command line are stored in the group-ADvsHC_glm.json file:
{
"DesignMatrix": "1 + age + sex + group"
"StringFormatTSV": "%s %f %f"
"Contrast": "group"
"ClusterThreshold": 0.001
}
The results of the group comparison between AD and HC are given by the group-ADvsHC_AD-lt-HC_measure-ct_fwhm-20_correctedPValue.jpg file and is illustrated as follows:
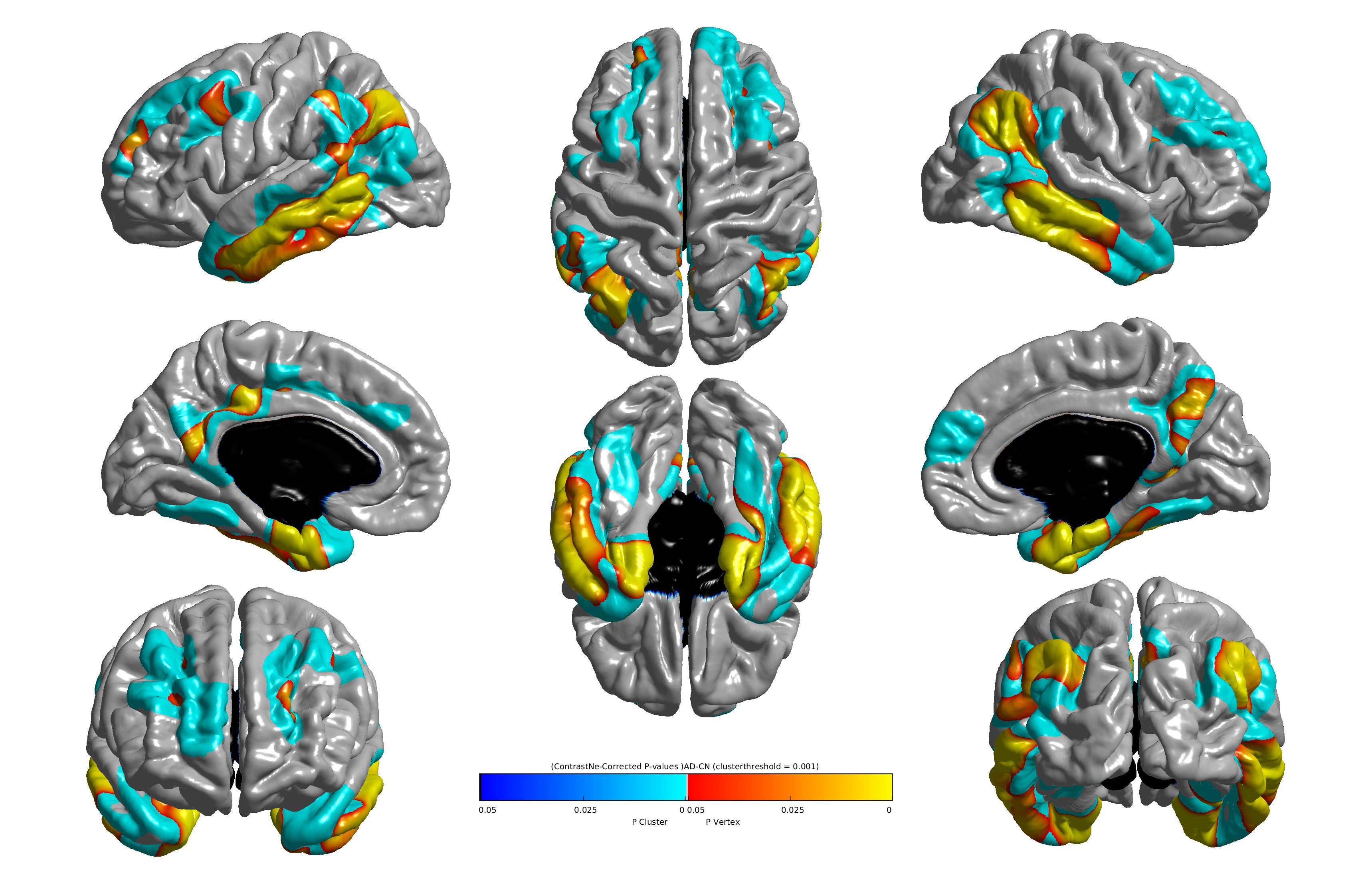
The blue area corresponds to the vertex-based corrected p-value and the yellow area represents the cluster-based corrected p-value.
Correlation analysis¶
Let's now assume that you are interested in knowing whether cortical thickness is correlated with age using the same population as above, namely ADvsHC_participants.tsv.
The contrast will become age and we will choose correlation instead of group_comparison.
The command line is simply:
clinica run statistics-surface caps_directory ADvsHC t1-freesurfer correlation ADvsHC_participants.tsv age -c group -c sex
Describing this pipeline in your paper¶
Example of paragraph (group comparison):
These results have been obtained using the statistics-surface command of Clinica [Routier et al., 2021].
More precisely, a point-wise, vertex-to-vertex model based on the Matlab SurfStat toolbox (http://www.math.mcgill.ca/keith/surfstat/) was used to conduct a group comparison of whole brain cortical thickness.
The data were smoothed using a Gaussian kernel with a full width at half maximum (FWHM) set to <FWHM> mm.
The general linear model was used to control for the effect of <covariate_1>, ... and <covariate_N>.
Statistics were corrected for multiple comparisons using the random field theory for non-isotropic images [Worsley et al., 1999].
A statistical threshold of P < <ClusterThreshold> was first applied (height threshold).
An extent threshold of P < 0.05 corrected for multiple comparisons was then applied at the cluster level.
Tip
Easily access the papers cited on this page on Zotero.
Support¶
- You can use the Clinica Google Group to ask for help!
- Report an issue on GitHub.
(Advanced) Specifying what surface data to use¶
If you run the help command line clinica run statistics-surface -h, you will find two optional flags:
--feature_type FEATURE_TYPEallows you to decide what feature type to take for your analysis. If it iscortical_thickness(default value), the thickness file for each hemisphere and each subject and session of the tsv file will be used. Keep in mind that those thickness files are generated using thet1-freesurferpipeline, so be sure to have run it before using it! Other directly-implemented solutions are present but they are not yet released.-
The other flag
--custom_file CUSTOM_FILEallows you to specify yourself what file should be taken in theCAPS/subjectsdirectory.CUSTOM_FILEis a string describing the folder hierarchy to find the file. For instance, let's say we want to manually indicate to use the cortical thickness. Here is the generic link to the surface data files:CAPS/subjects/sub-*/ses-M*/t1/freesurfer_cross_sectional/sub-*_ses-M*/surf/*h.thickness.fwhm*.fsaverage.mghExample:
CAPS/subjects/sub-ADNI011S4075/ses-M000/t1/freesurfer_cross_sectional/sub-ADNI011S4075_ses-M000/surf/lh.thickness.fwhm15.fsaverage.mghNote that the file must be in the
CAPS/subjectsdirectory. So myCUSTOM_STRINGmust only describe the path starting after thesubjectsfolder. So now, we just need to replace the*by the correct keywords, in order for the pipeline to catch the correct filenames.@subjectis the subject,@sessionthe session,@hemithe hemisphere,@fwhmthe full width at half maximum. All those variables are already known, you just need to indicate where they are in the filename!As a result, we will get for
CUSTOM_FILEof cortical thickness:@subject/@session/t1/freesurfer_cross_sectional/@subject_@session/surf/@hemi.thickness.fwhm@fwhm.fsaverage.mghYou will finally need to define the name your surface feature
--feature_label FEATURE_LABEL. It will appear in the_measure-<FEATURE_LABEL>of the output files once the pipeline has run.
Note that --custom_file and --feature_type cannot be combined.
Appendix¶
- For more information about SurfStat, please check here.
- For more information about the GLM, please check here.
- The cortical thickness map is obtained from the FreeSurfer segmentation.
More precisely, it corresponds to the subject’s map normalized onto FSAverage and smoothed using a Gaussian kernel FWHM of
<fwhm>mm (thesurf/?h.thickness.fwhm<fwhm>.fsaverage.mghfiles).